pkCSM - pharmacokinetics
pkCSM: predicting small-molecule pharmacokinetic properties using graph-based signatures
Douglas E. V. Pires, Tom L. Blundell, David B. AscherJournal of Medicinal Chemistry, 58 (9), p. 4066–4072, 2015.


Abstract
Modern high throughput drug discovery approaches have increased the numbers of lead compounds
being identified, and in shorter time frames than traditional medicinal chemistry, however many of these promising
compounds often fail because of unsatisfactory ADMET properties. In silico screening approaches help to reduce these
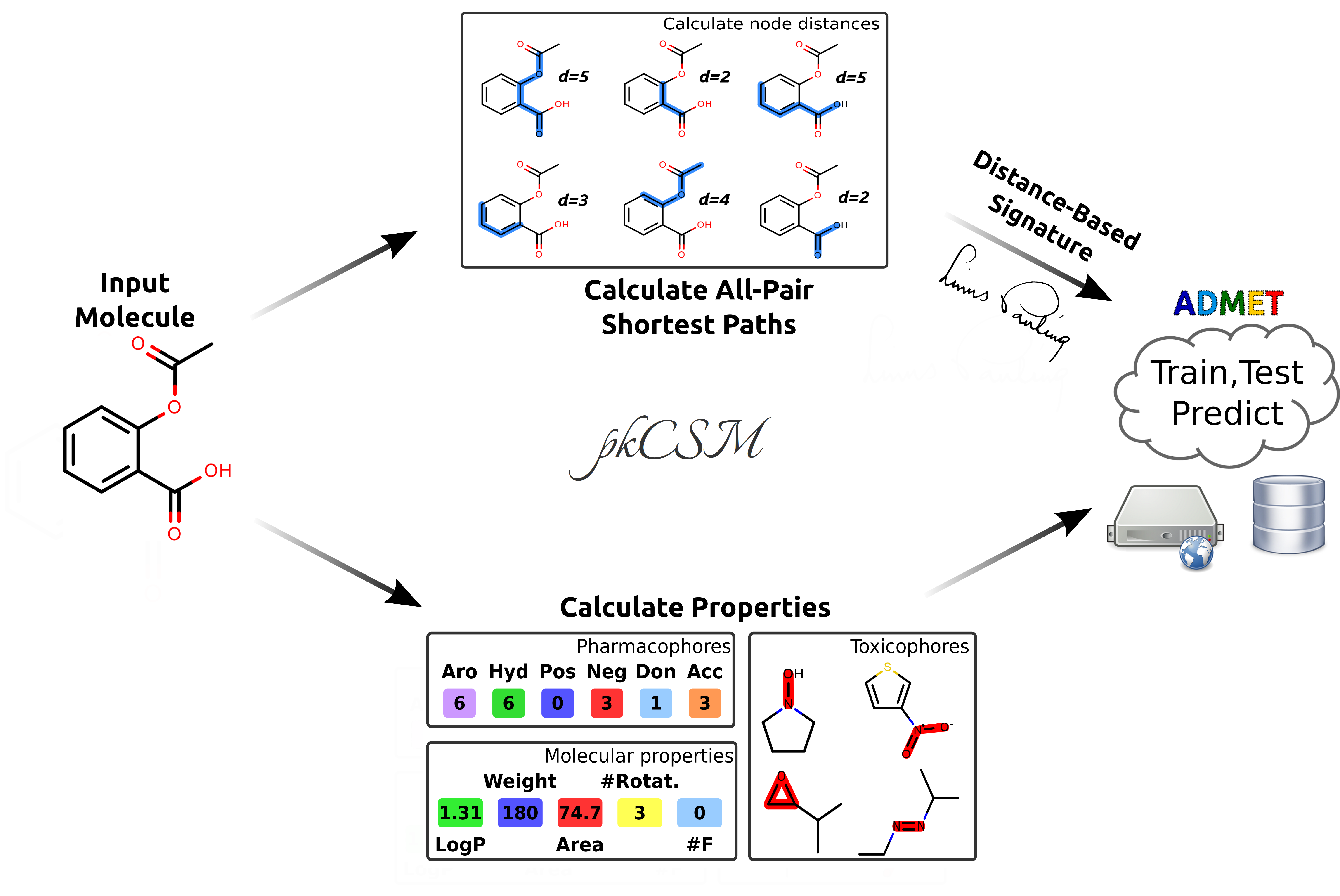
risks. Here we propose a novel approach to the prediction of pharmacokinetic properties, called pkCSM, which relies
on graph-based signatures. These encode distance patterns between atoms and are used to represent the small molecule
and to train predictive models.
The pkCSM signatures were successfully used across five main different pharmacokinetic properties classes
to develop predictive regression and classification models. We show that pkCSM performs as well or better across
different pharmacokinetic properties than other freely available methods.
Here we present a web server to provide an integrated freely available platform to rapidly screen multiple
pharmacokinetic properties.