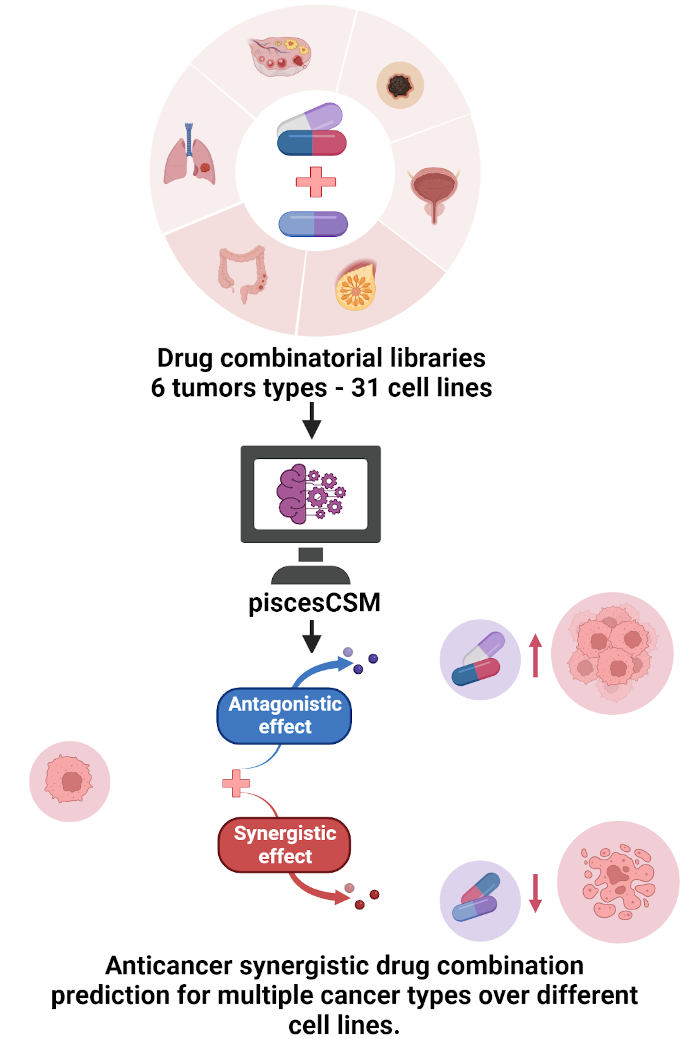
piscesCSM accurate prediction of anticancer synergistic drug combinations
Raghad Aljarf, Yoochan Myung, Carlos H.M. Rodrigues, Douglas E.V. Pires & David B. Ascher
Abstract: While drug combination therapies are of great importance in cancer treatment, novel synergistic combination identification has been specifically challenging; owing to the size of the combinatorial search space. Nevertheless, computational methods have emerged as a faster and cost-efficient alternative for prioritizing drug combinations prediction in large-scale combinatorial screening libraries. This study proposes a novel predictive tool called piscesCSM. piscesCSM leverages the graph-based signature representation of a small molecule's chemical structure to accurately predict drug combinations with favorable anticancer synergistic effects against one or multiple cancer cell lines in a machine learning framework. Leveraging these insights, we developed a generic predictive model to guide the prediction of anticancer synergistic drug combinations in at least 31 cell lines. It achieved an area under the receiver operating characteristic curve (AUC) of up to 0.89 and 0.87 on independent non-redundant blind tests. We compared piscesCSM with the state-of-the-art approaches on a large-scale oncology screen data set. The leave-one-out validations results demonstrated that our predictive models outperformed competitive methods. Furthermore, our model was superior to alternative approaches by more than 16% predictive accuracy on an independent test set published by the pharmaceutical enterprise AstraZeneca.To provide a simple and integrated platform to rapidly screen potential candidate pairs with favorable synergistic anticancer effects, we made piscesCSM freely available online. We believe that our predictive tool will provide a valuable resource for optimizing and augmenting combinatorial screening libraries to identify effective and safe synergistic anticancer drug combinations.
searchRun predictionhouseThe standalone version of piscesCSM is available for download.