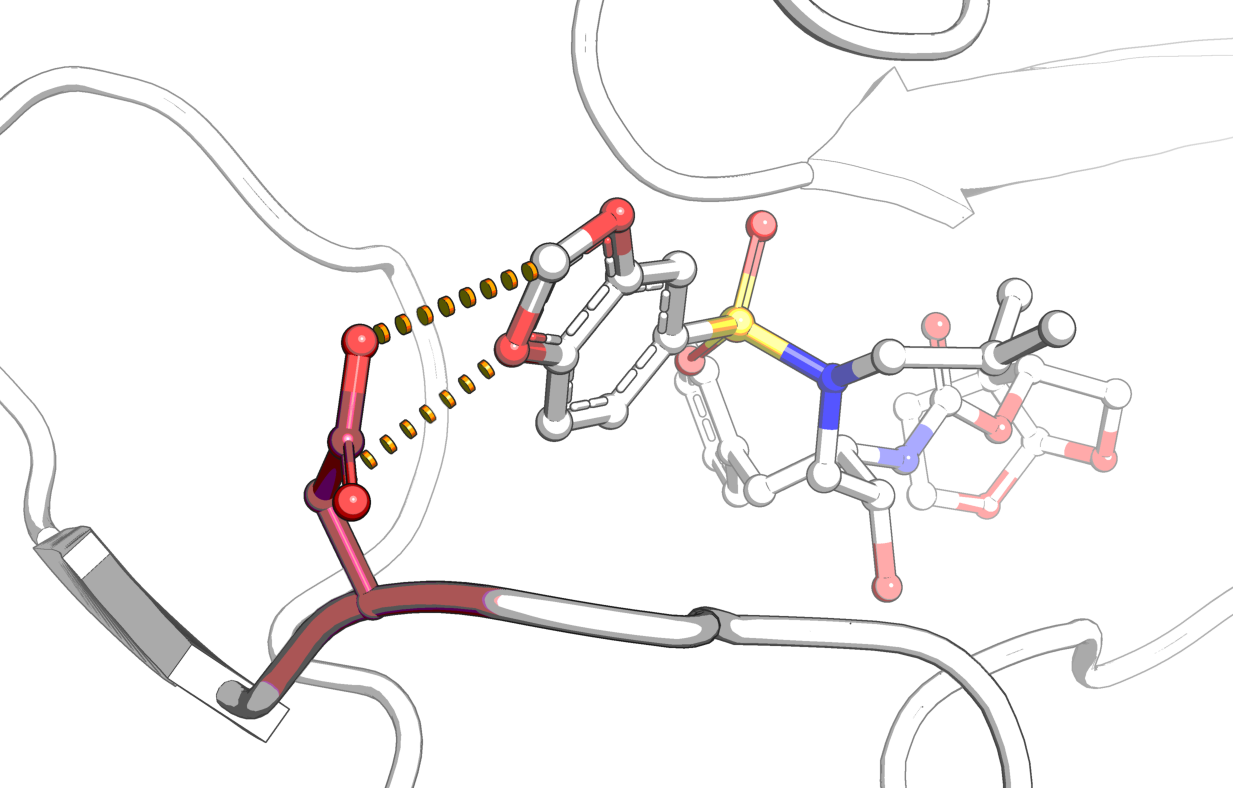
mCSM-lig
mCSM-lig: quantifying the effects of mutations on protein-ligand affinity in genetic disease and the emergence of drug resistance
Douglas E. V. Pires, Tom L. Blundell, David B. AscherScientific Reports, 6(29575), 2016


Abstract
The emergence of drug resistance is a growing problem limiting the effectiveness of many medicines. The ability to predict how a mutation affects
ligand binding is an essential step in understanding, anticipating and improving the design of new treatments for drug resistance, understanding
genetic diseases and in engineering proteins with new ligand specificities.
Here we present mCSM-lig, a structure-guided in silico approach for directly quantifying the effects of single-point missense mutations on
affinities of small molecules for proteins. mCSM-lig uses graph-based signatures to train a predictive model using a representative set of
protein-ligand complexes from the Platinum database. We show our method presented very good correlation with experimental data (up to ρ=0.67),
and was effective in predicting a range of resistance mutations, explaining Mendelian disease mutations, and in guiding protein design.