How to use DPD-Cancer
A comprehensive guide to molecular anti-cancer prediction and analysis.
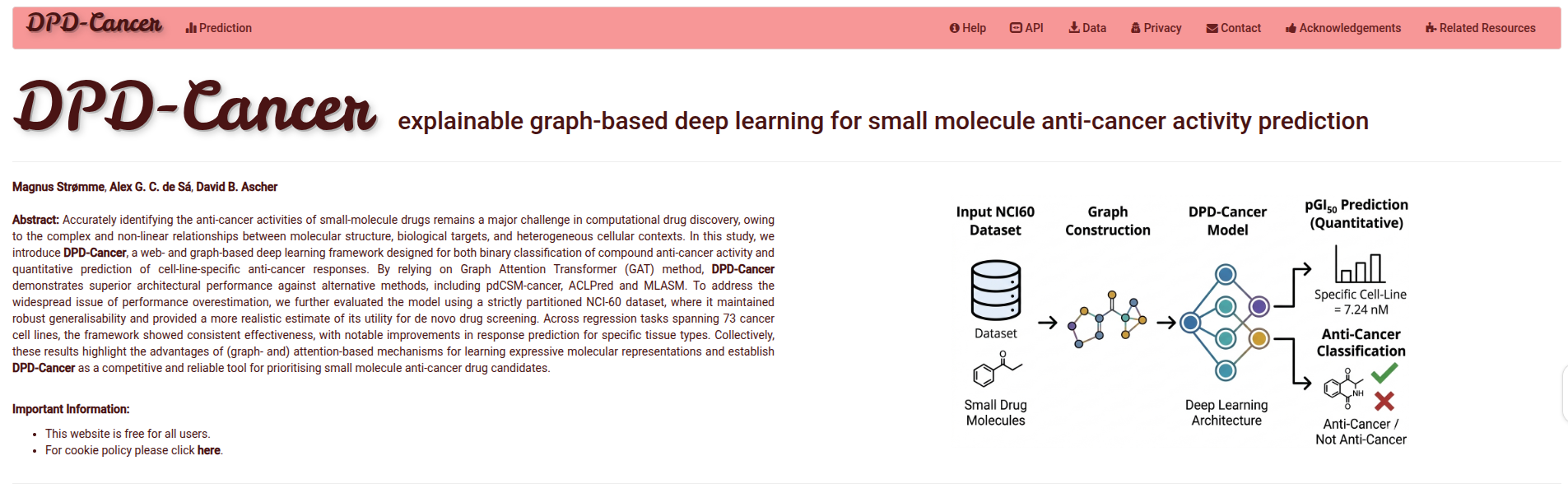
System Overview
DPD-Cancer leverages Graph Neural Networks (GNN) and molecular descriptors to provide high-fidelity predictions for 73 cell-line anti-cancer predictors based on pGI50% and one general anti-cancer predictor classifier.
- Prediction: Submit jobs via the web interface or locally via API.
- Transparency: Access documentation or download the full Experimental Datasets.
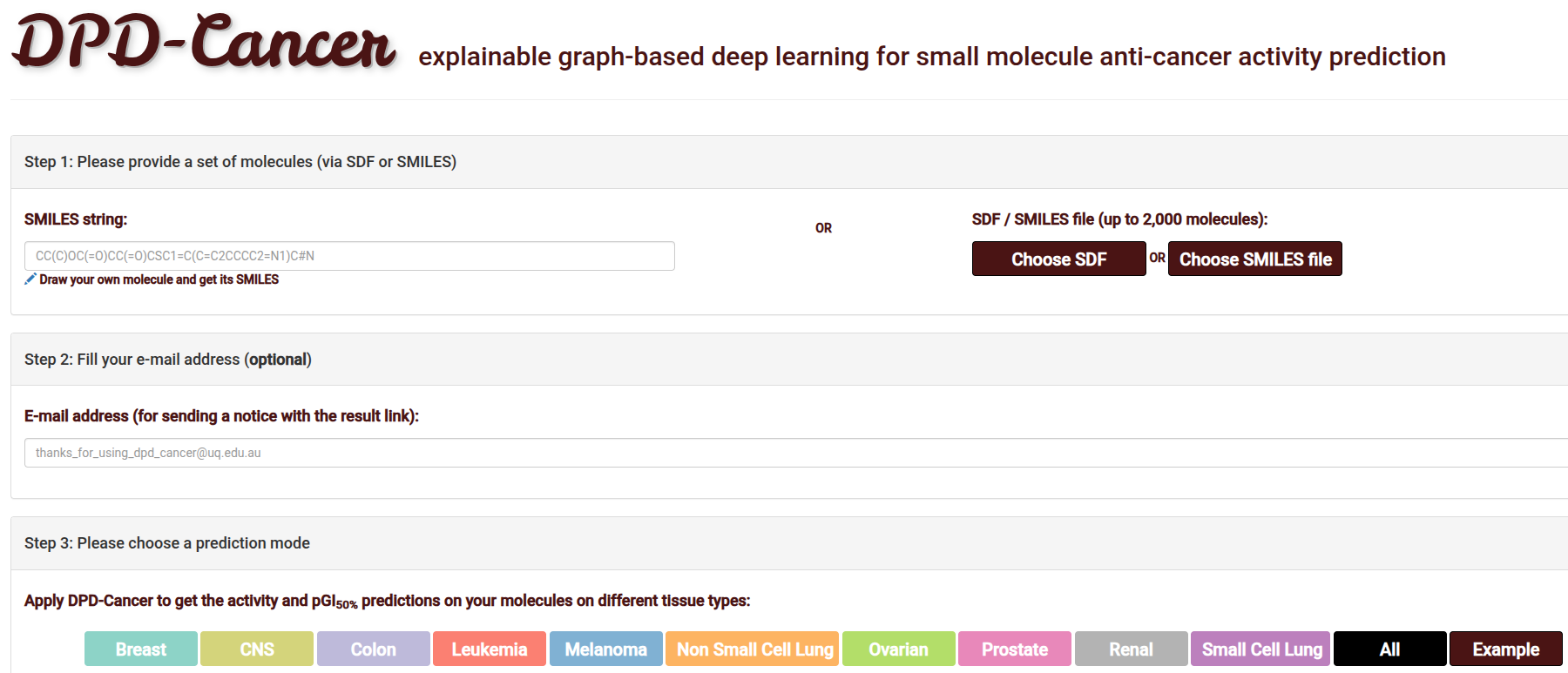
Job Submission
Flexible input options for individual or batch molecular analysis:
- File Upload: Supports SDF or SMILES formats.
- Direct Entry: Paste a SMILES string or use the JSME Editor to sketch structures.
- Execution Mode: Select specific cancer cell lines (e.g., Leukemia, Renal, CNS) or run "All".
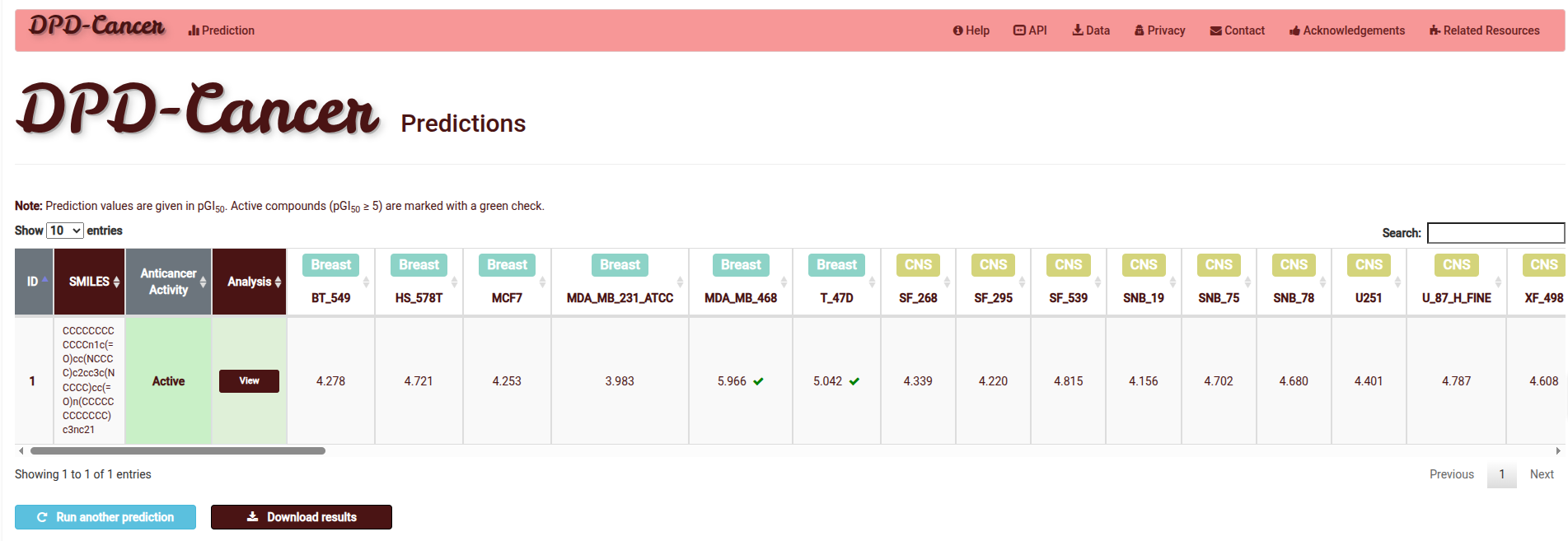
Predictive Performance and Bioactivity Mapping
The Results Page provides a quantitative assessment of molecular potency through pGI50% modelling, supplemented by instant bioactivity indicators:
- Potency Quantification: Bioactivity is classified across all cell lines and cancer types, derived directly from the underlying pGI50% thresholds.
- Global Classificatio: Paste a SMILES string or use the JSME Editor to sketch structures.
- Contextual Tooltips: Hover over specific data points to trigger a tooltip indicating if a molecule is "active" or "inactive" for a particular cell line or cancer type.
- Cell-Line Specificity: Analyse performance across diverse categories—including Breast, CNS, Leukemia, and Melanoma—to identify selective anti-cancer properties.
- Actionable Insights: Use the combined regression and activity data to prioritize candidates for further experimental validation or substructure importance analysis.
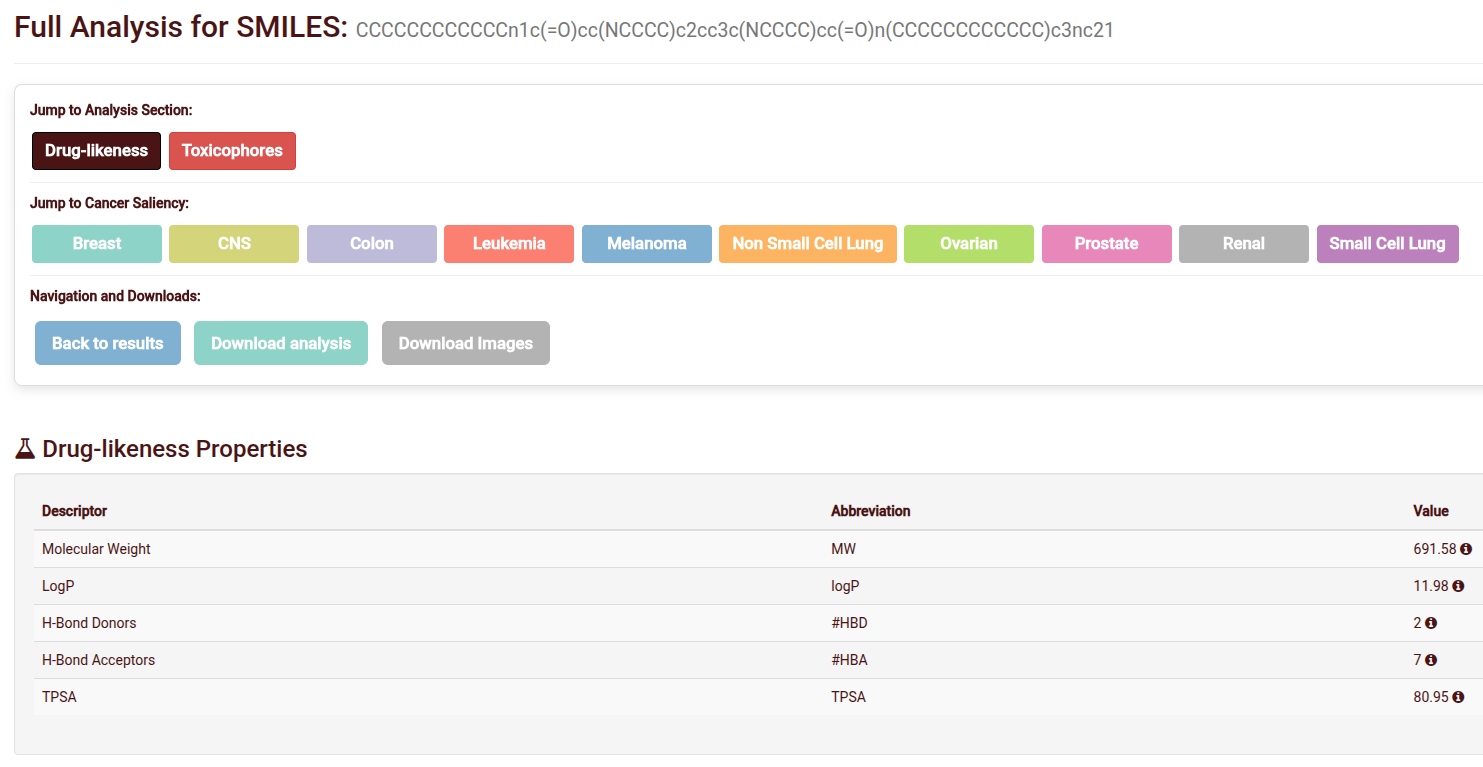
Deep Dive Analysis
The Analysis Page provides a granular look at chemical safety and model reasoning:
- Molecular properties: Calculate common molecular properties and link them with drug likeness.
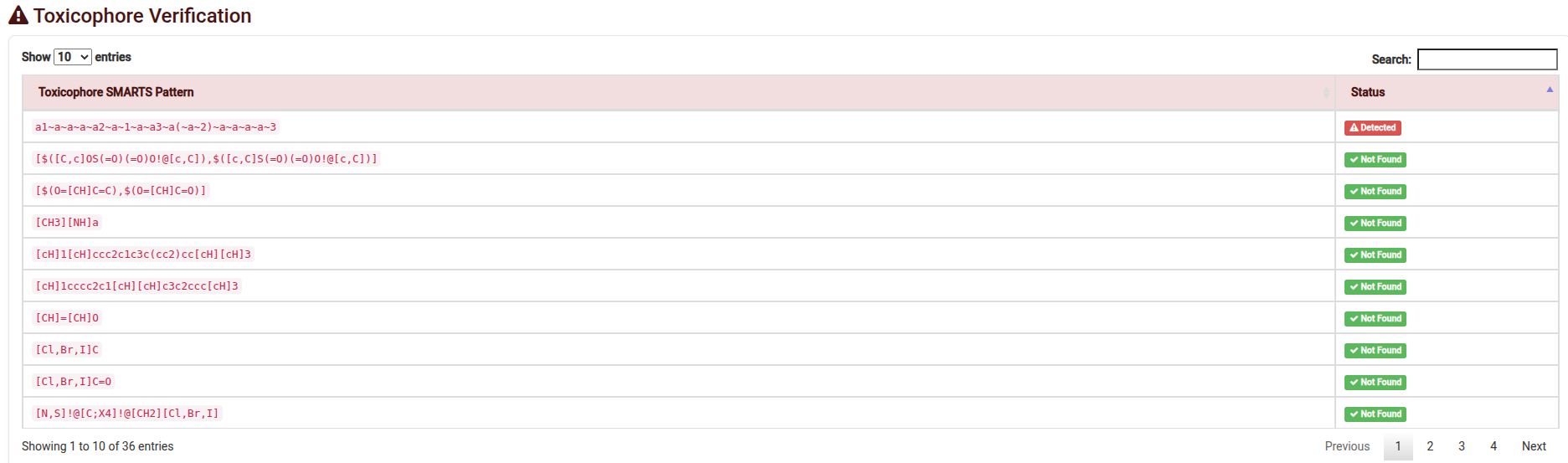
- Toxicophore Search: Automatically flags substructures associated with genomic toxicity.
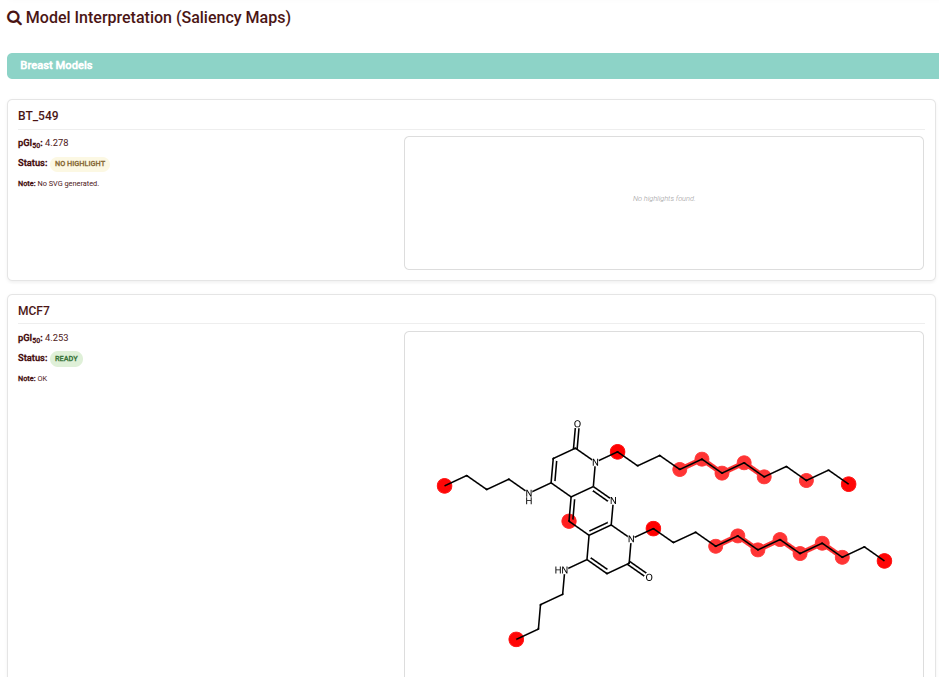
- Feature Importance: Visualizes the weight of molecular substructures in the model's decision-making process.
- Reporting: All images and tabular data are available for download.
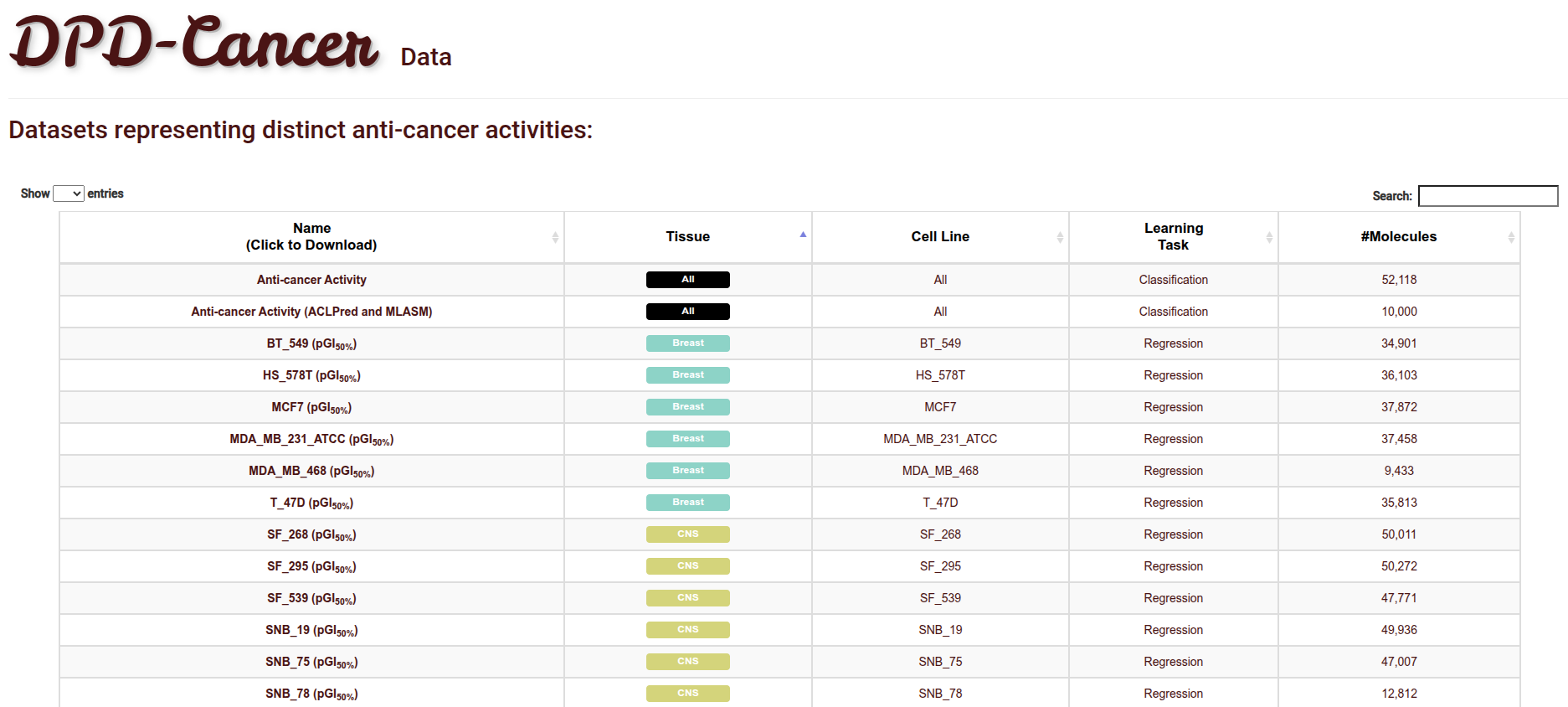
Data Availability
The Data Page makes the whole molecular data available not only generally (i.e., considering all cell lines and cancer lines for molecular anti-cancer classification), but also per each combination of cancer type and cell line.