CSM-Toxin provides a comprehensive suite for rapid identification of toxin proteins.
Running predictions
Input
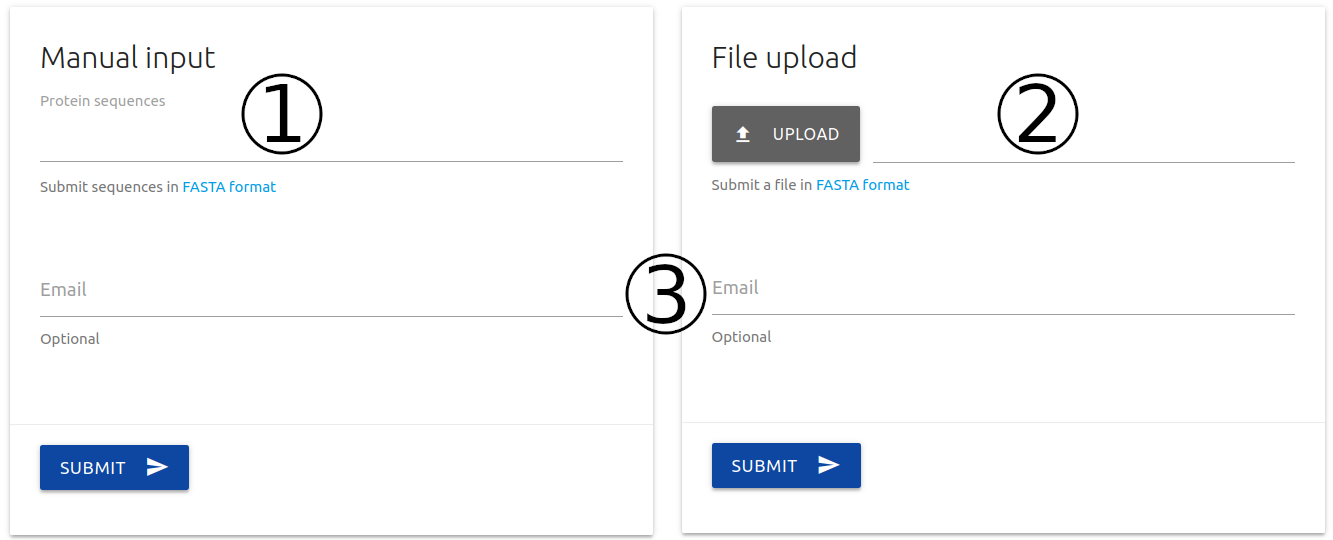
The input page can be accessed from the home page or via the top menu Run. Here, two options are available:
-
Manual - Submitting one or multiple (max 10) protein sequences in FASTA format (1)
-
File upload - Submitting a file in fasta format with a list of protein sequences (2)
For both types of submissions, users may provide an email (3) which will be used to send the results once they are ready.
Results
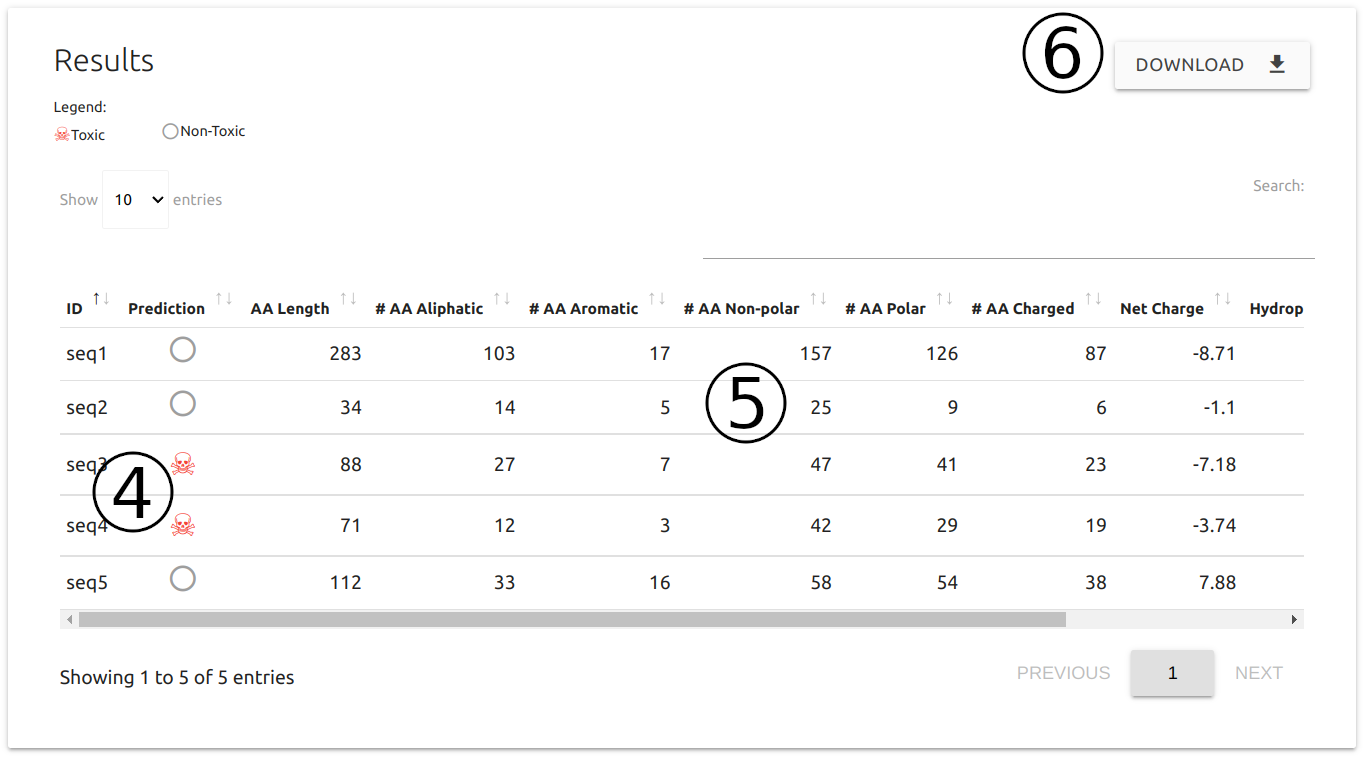
Results are displayed in table format:
- Predictions for each protein sequence are shown (4) alongside with general physicochemical details (5)
- All results can be downloaded as a comma separated file (CSV) (6)
Contact us
In case you experience any troube using CSM-Toxin or if you have any suggestions or comments, please do not hesitate in cantacting us either via email or via our Group website.
If your are contacting regarding a job submission, please include details such as input information and the job identifier