1.Submission Page
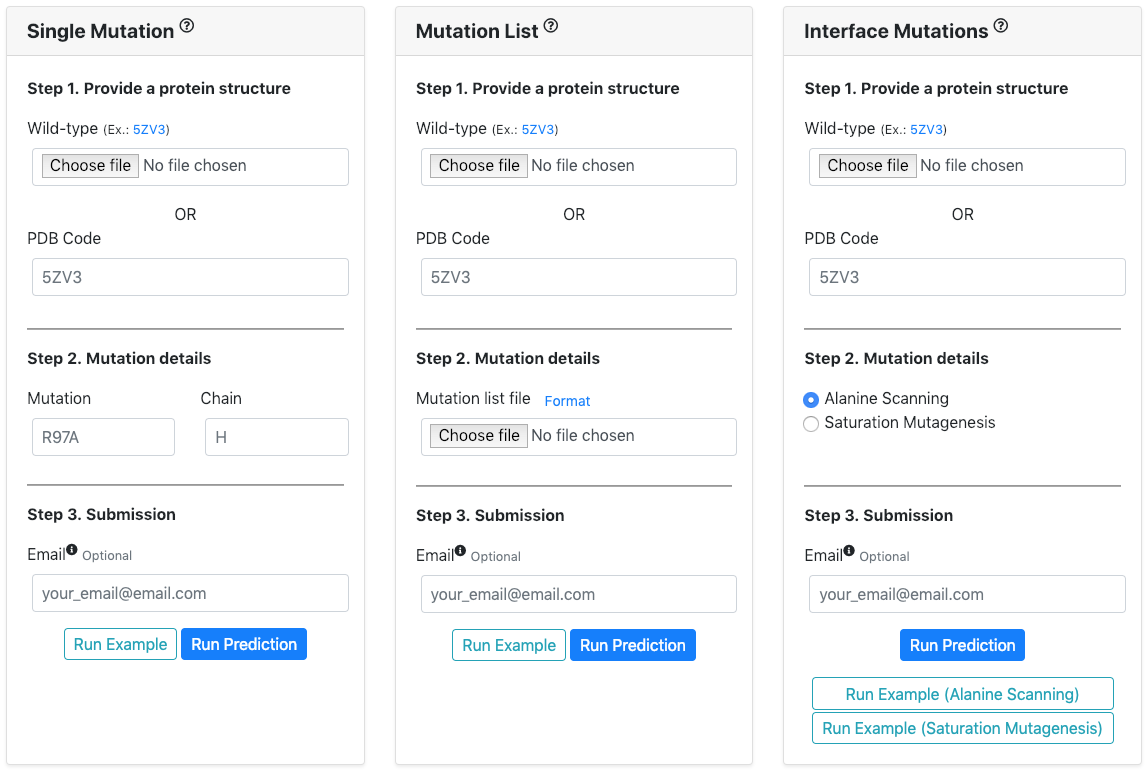
Users can run three types of prediction.
Single Prediction gives single ∆∆G for given antibody-antigen structure and information about single mutation.
Multiple Prediction allows to predict ∆∆G for multiple mutations with list of mutations.
Also, users are able to conduct Alanine Scanning or Saturation Mutagenesis for antibody-antigen interface residues.
2.Result Page- Single Prediction (1/2)
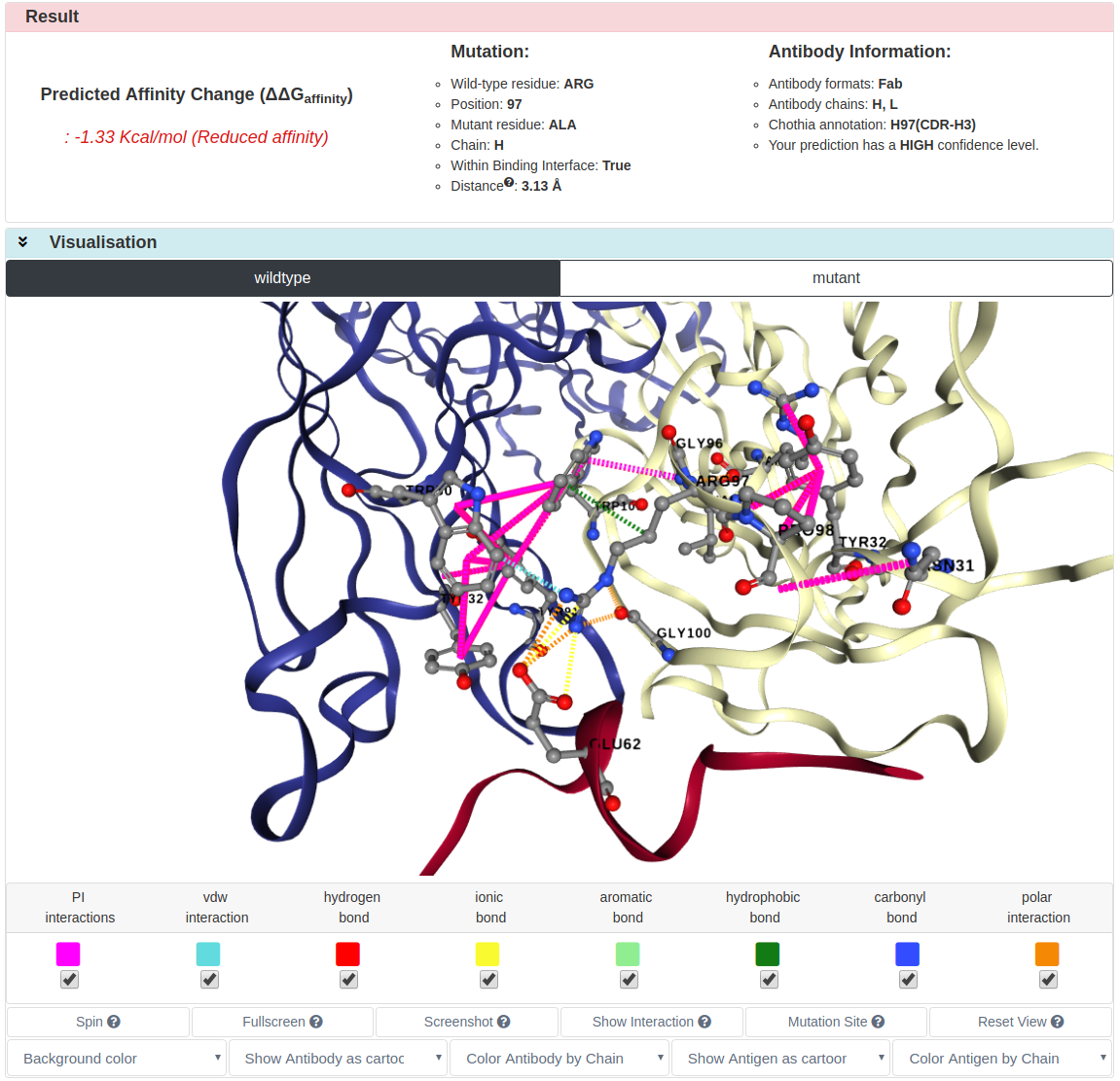
The result page of Single Prediction is divided into three sections.
The Result section gives predicted ∆∆G, mutation information and antibody
information.
The Visualisation part shows inter-/intramolecular interactions of wild-type and mutant
structures. Using control panel, users are able to turn on/off each interaction and change the representation of antibody-antigen structure.
3.Result Page- Single Prediction (2/2)
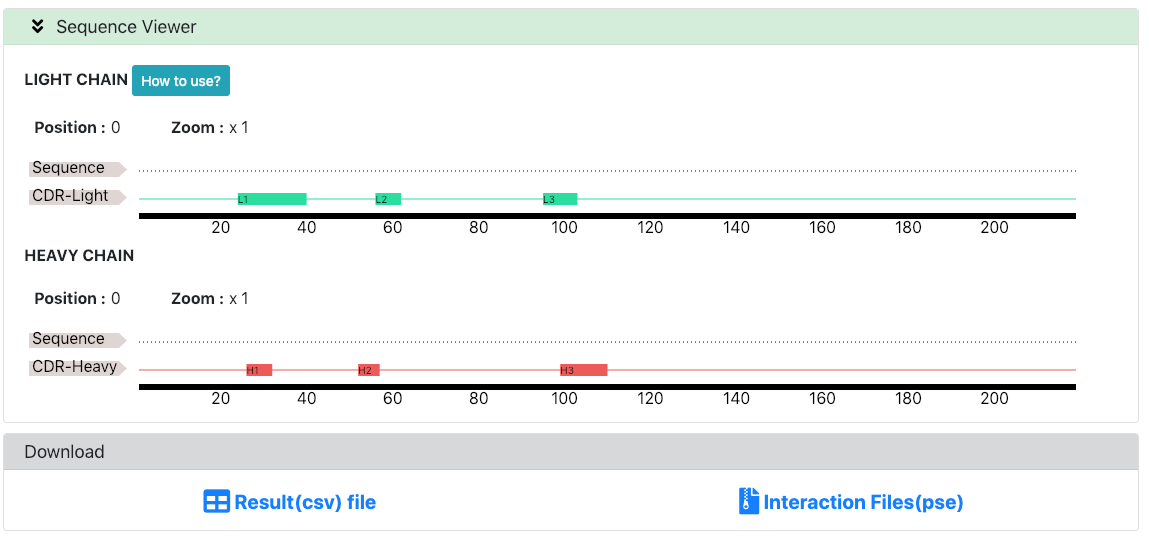
The antibody sequence and its Chothia annotation are shown in the Sequence Viewer.
Users are able to download results and interaction files through the Donwload section.
4.Result Page- Multiple Mutation Prediction (1/2)
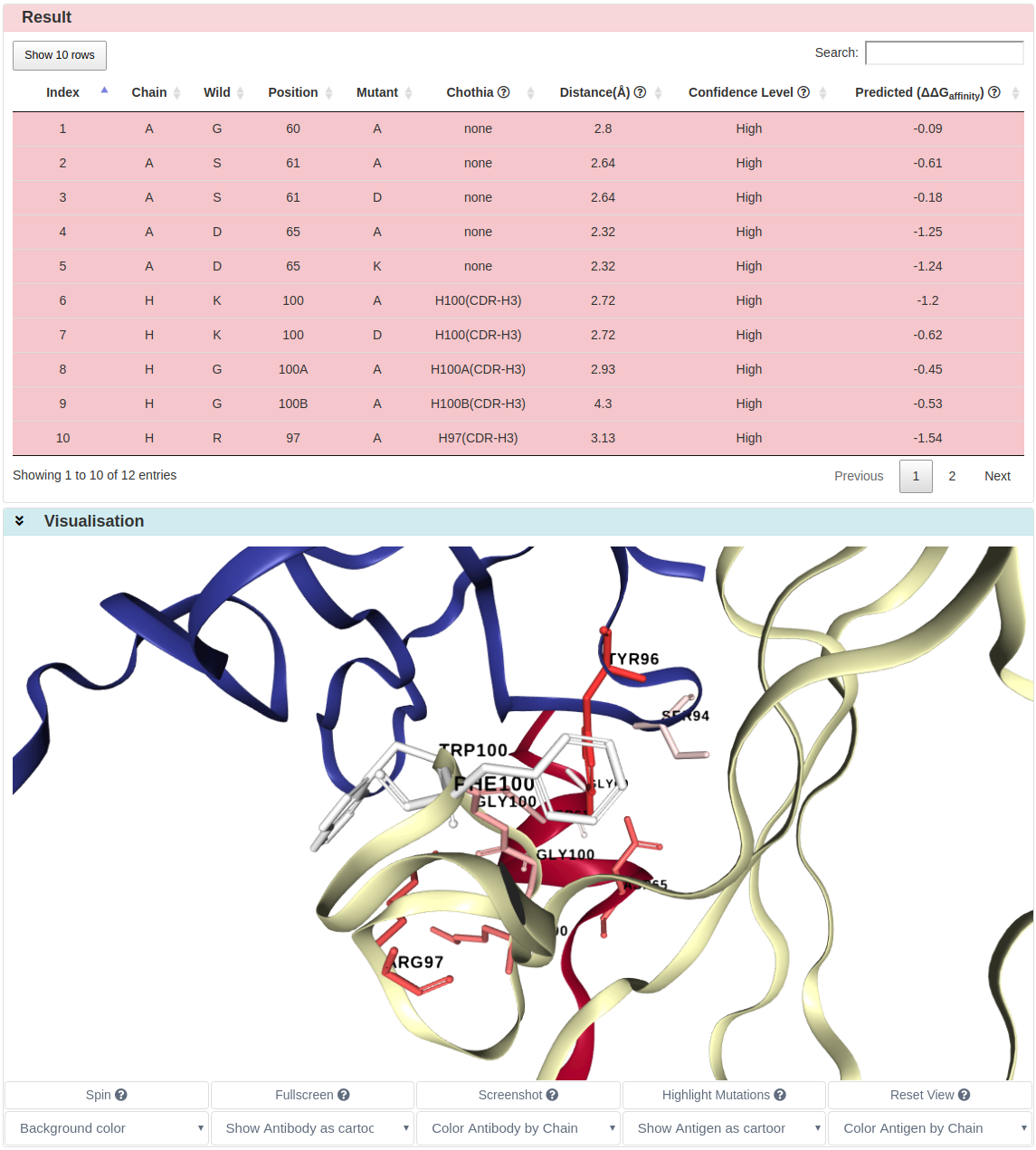
The predicted ∆∆G and corresponding information are shown in the Result section.
The antibody-antigen complex can be coloured by average ∆∆G if any of mutations occurs more than once at the same position.
5.Result Page- Multiple Mutation Prediction (2/2)
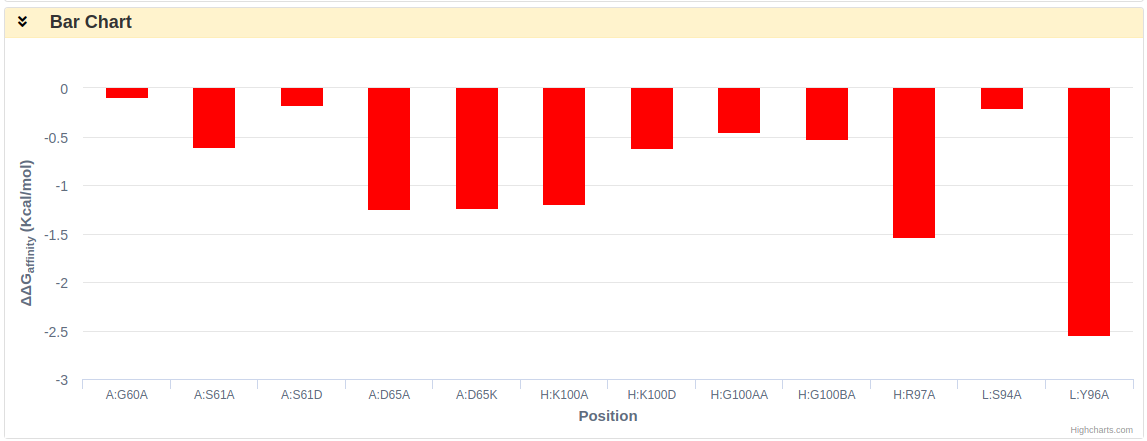
All the predicted ∆∆G are plotted in the Bar Chart.
The result table and Antibody-Antigen complex which of B-factor is replaced with average ∆∆G are downloadable through the Download section.
6.Result Page- Alanine Scanning
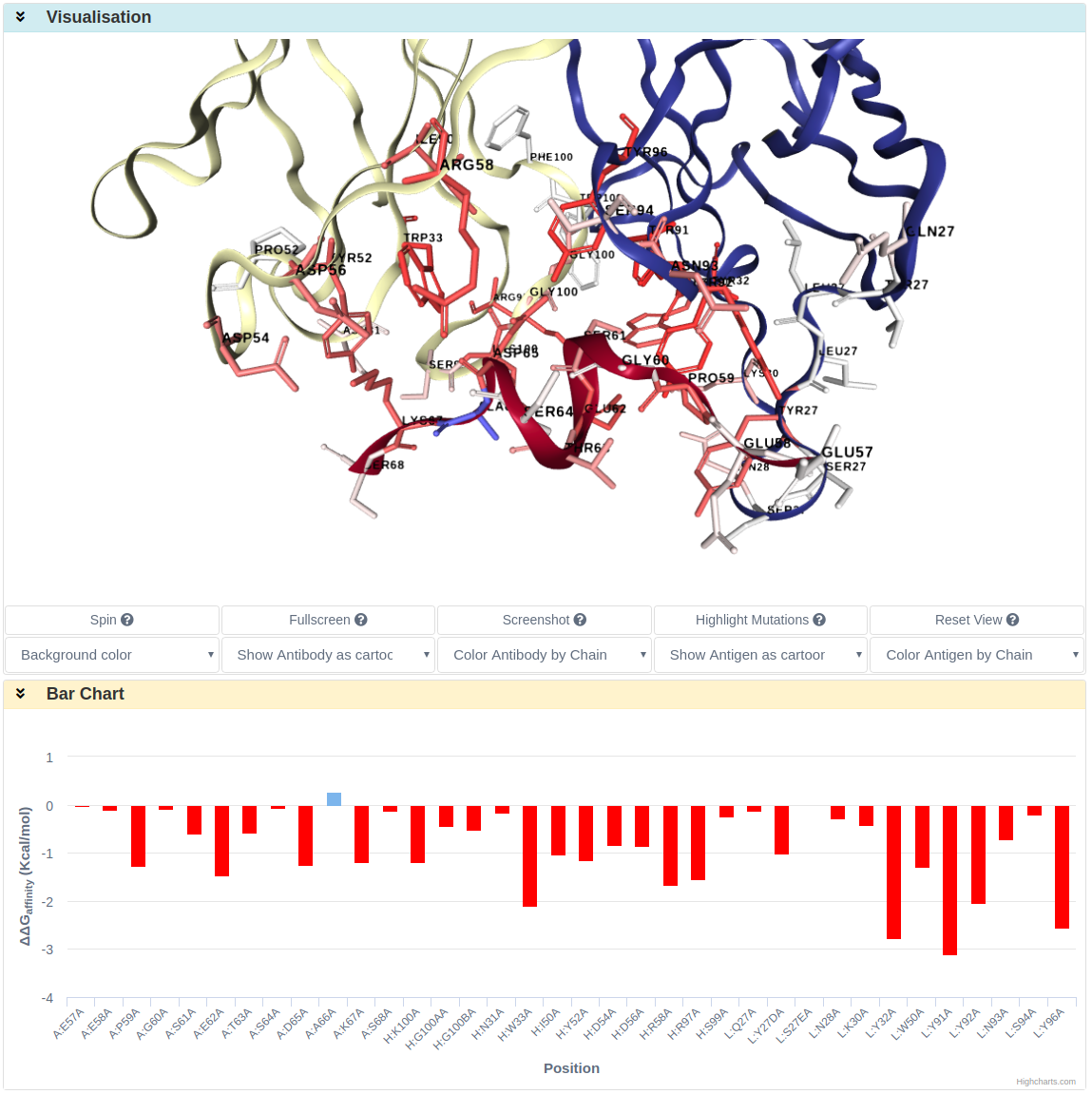
Alanine Scanning gives predicted ∆∆G for antibody-antigen interface residues. Same as the previous mutation list prediction, the result is shown in Result table, Visualisation and Bar Chart sections.
7.Result Page- Saturation Mutagenesis
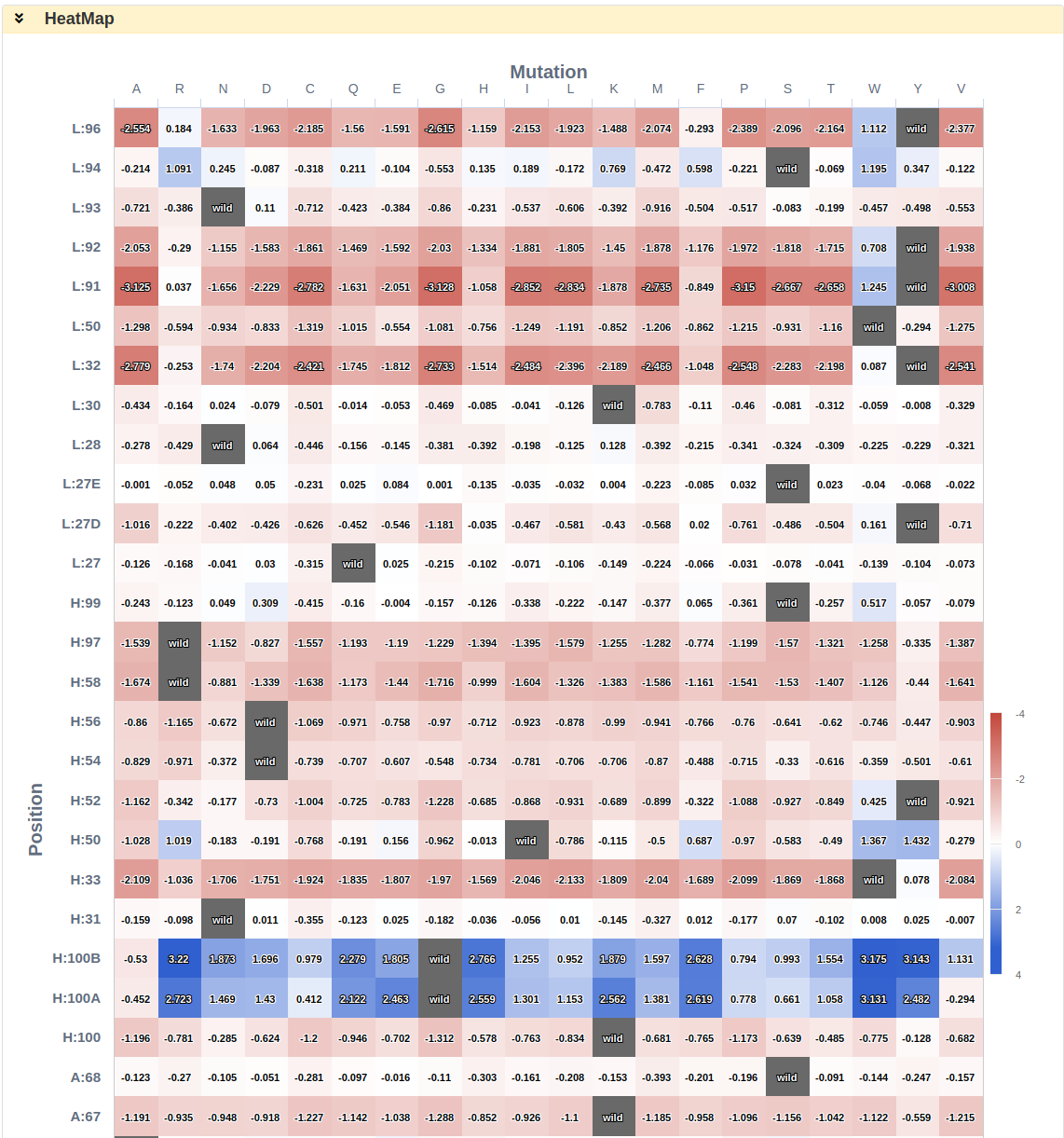
Apart from the Result and Visualisation sections, Saturation Mutagenesis shows a series of predicted ∆∆G for each of antibody-antigen interface reisdues using Heat Map.