1. Predict: Submission Page
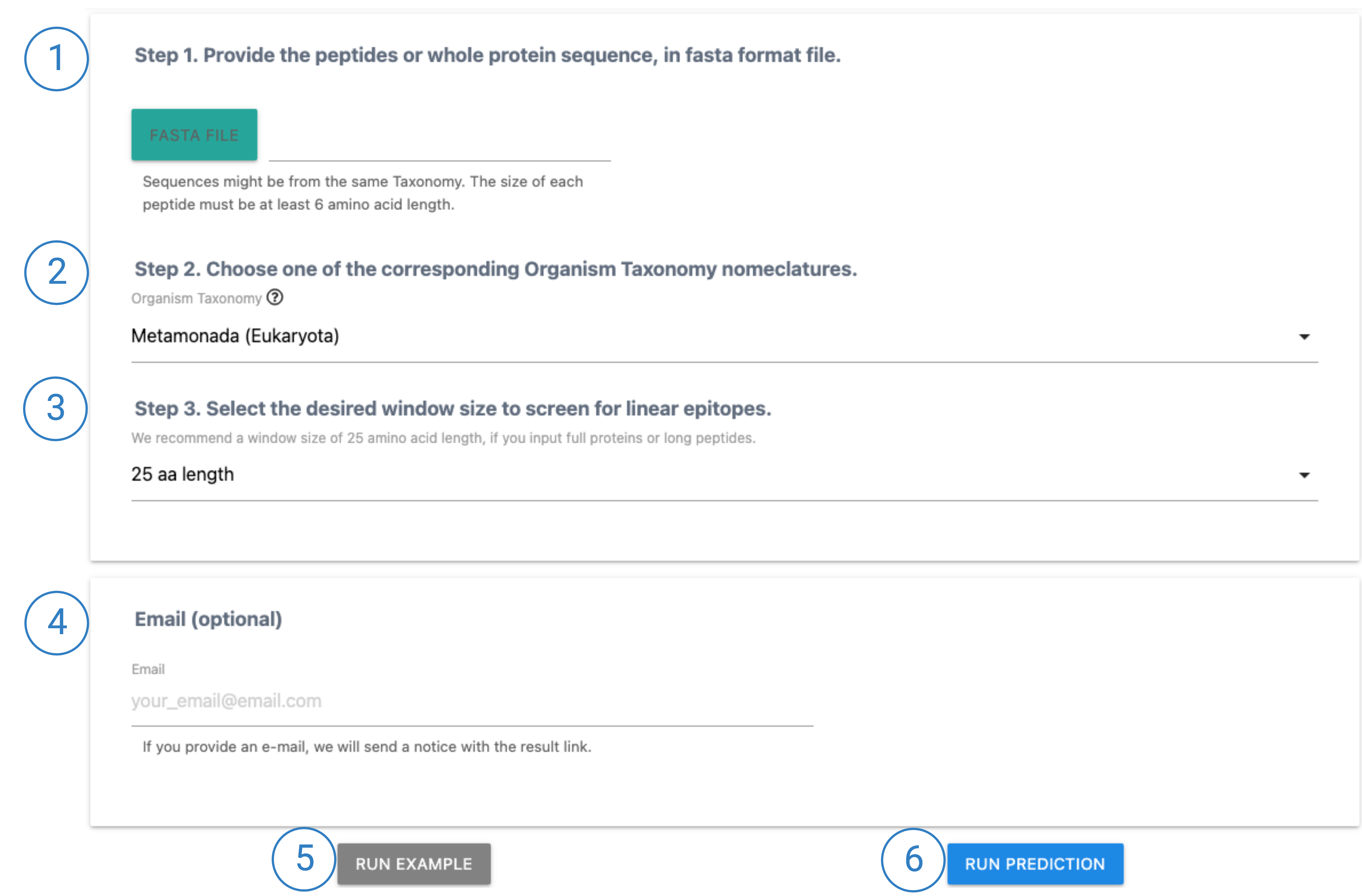
1. Multiple peptides or protein sequences can be provided as input in fasta format file. Each sequence should be placed in a different line and must have a header, starting with '>'. Also, sequences should be of the same Taxonomy group and be at least 6 amino acid in length.
2. Choose from the drop-down menu one of the 20 possible Organism Taxonomy nomeclatures that your sequences belong to.
3. Linear b-cell epitope sequences range (on average) from 6 to 25 amino acid length. Please select the window size you would like to set, so that our model can screen the given fasta file using this parameter. We recommend window = 25.
4. Provide user's email (optional) if one would like to be advised when the prediction is finished.
5. Instead, if user would like to view an example of Result's page, click 'Run Example' button.
6. Click RUN PREDICTION to submit the provided fasta file for prediction.
2. Result Page: Linear Epitope Prediction
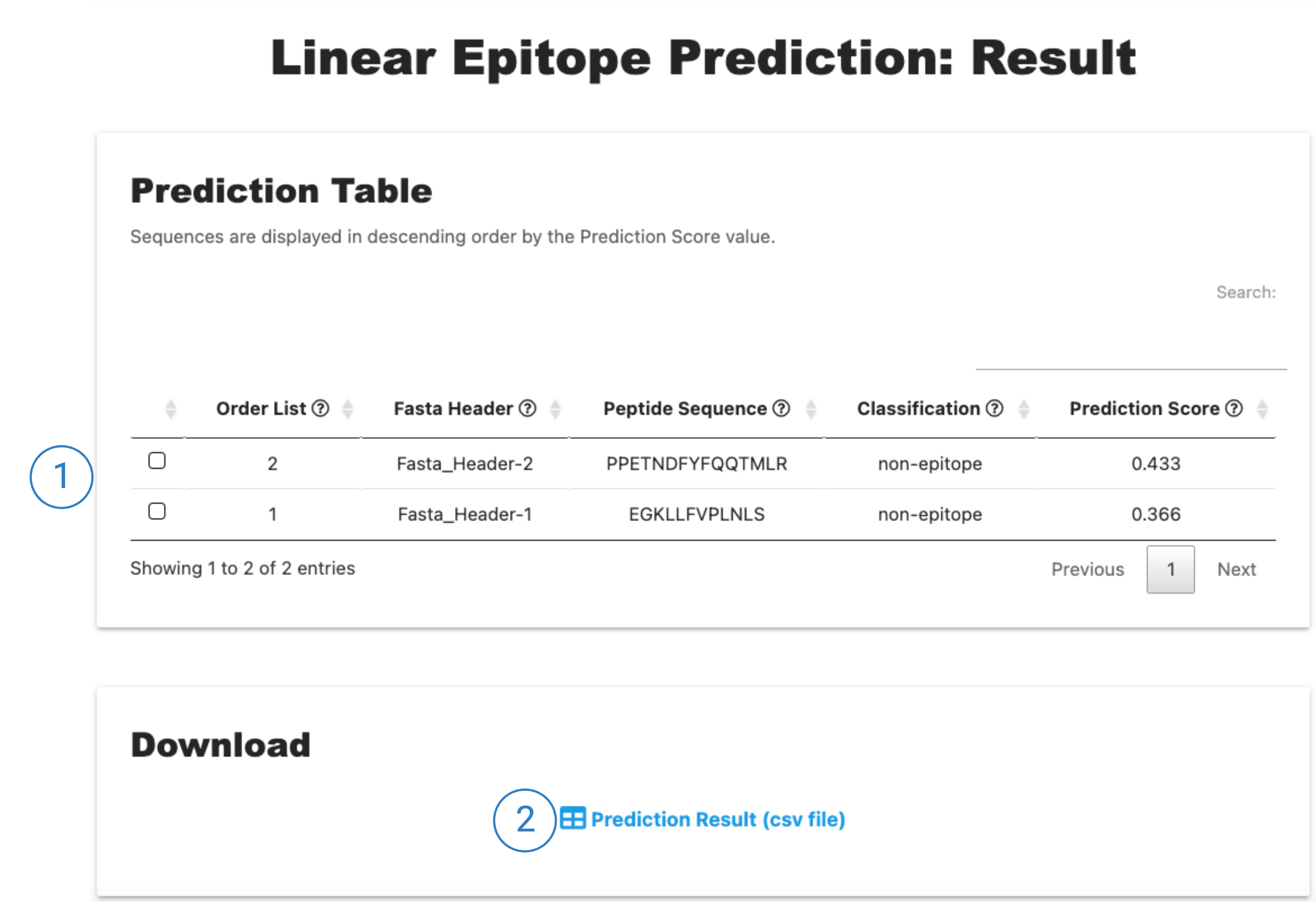
The result page is divided into two sections:
1. The Prediction Table section shows the output epitope prediction for each peptide sequence, of the Fasta file. The Order List column is a numerical list os peptides as observed in the Fasta file, followed by the Fasta Header, the Peptide Sequence (according to the window size informed), the Classification output (epitope means that the model predicted that peptide sequence as a linear epitope, or non-epitope) and finally the Prediction Score, showing the result probability score (0 to 1).
2. The Prediction table is available to be downloaded as a .csv file.