mCSM-metal : A deep learning resource to predict effect of mutations on metal ion binding.
Akshita Kumar, Ashar Malik, and David B. AscherWelcome to mCSM-metal
Metal ions play several major roles in proteins, including structural, regulatory, and enzymatic functions, making their binding essential for biological processes. While experimental methods for identifying metal-binding sites are resource-intensive and limited in scalability, recent advancements in large language models and protein embeddings have revolutionized computational approaches in this field. However, existing tools are still unable to address how residue-level metal-binding probabilities change under the effect of mutations. To fill this gap, mCSM-metal leverages state-of-the-art deep learning embeddings from ESMBind with our graph-based structural signatures to accurately predict the effects of single or multiple point mutations on the binding of seven essential ions (Zn²⁺, Ca²⁺, Mg²⁺, Mn²⁺, Fe³⁺, Co²⁺, Cu²⁺). This resource provides researchers with an intuitive platform to assess and visualize local and long range impacts of mutations on ion binding, offering new avenues for applications in structural biology, disease modelling, and protein engineering.
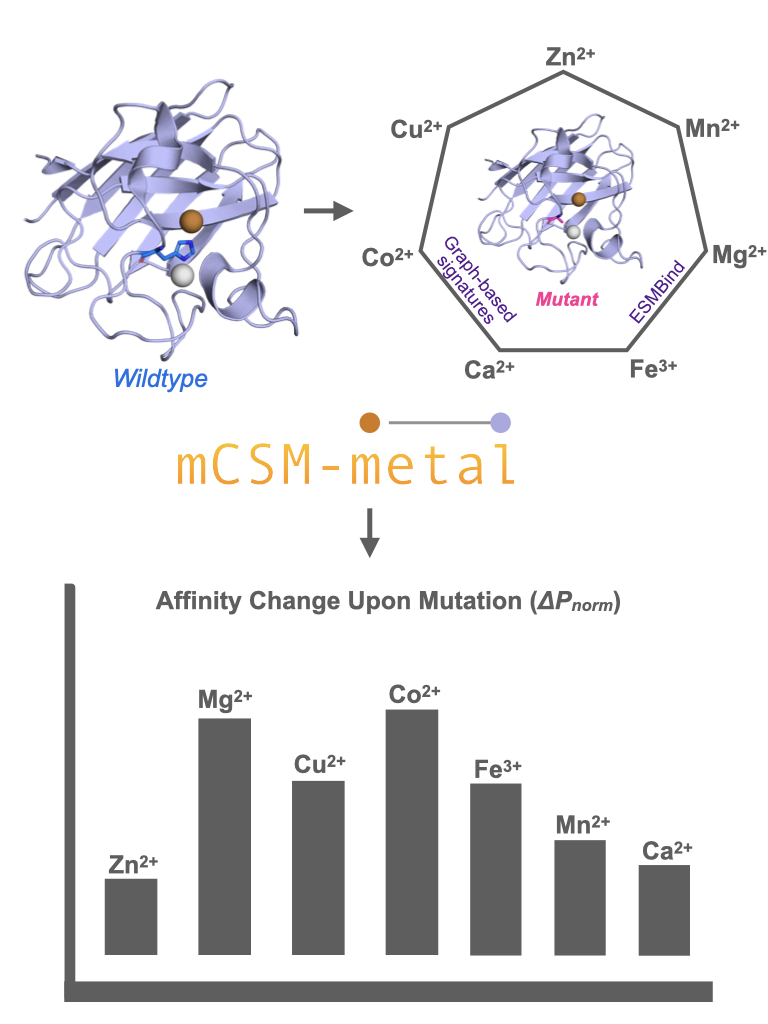
From single change to binding fate - see what mCSM-metal reveals!