Deep homology using high-dimensional latent space embeddings
Ashar J. Malik, and David B. Ascher
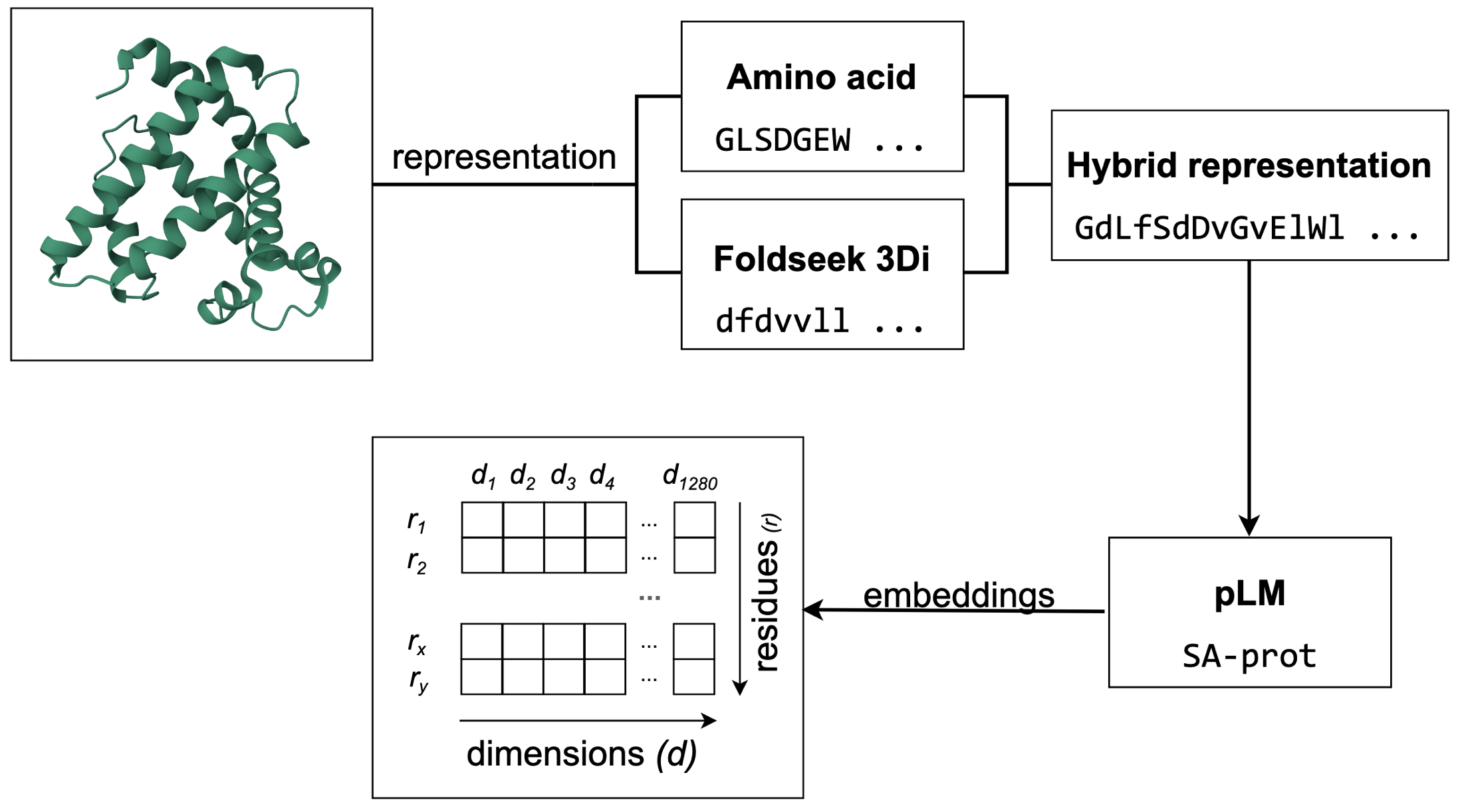
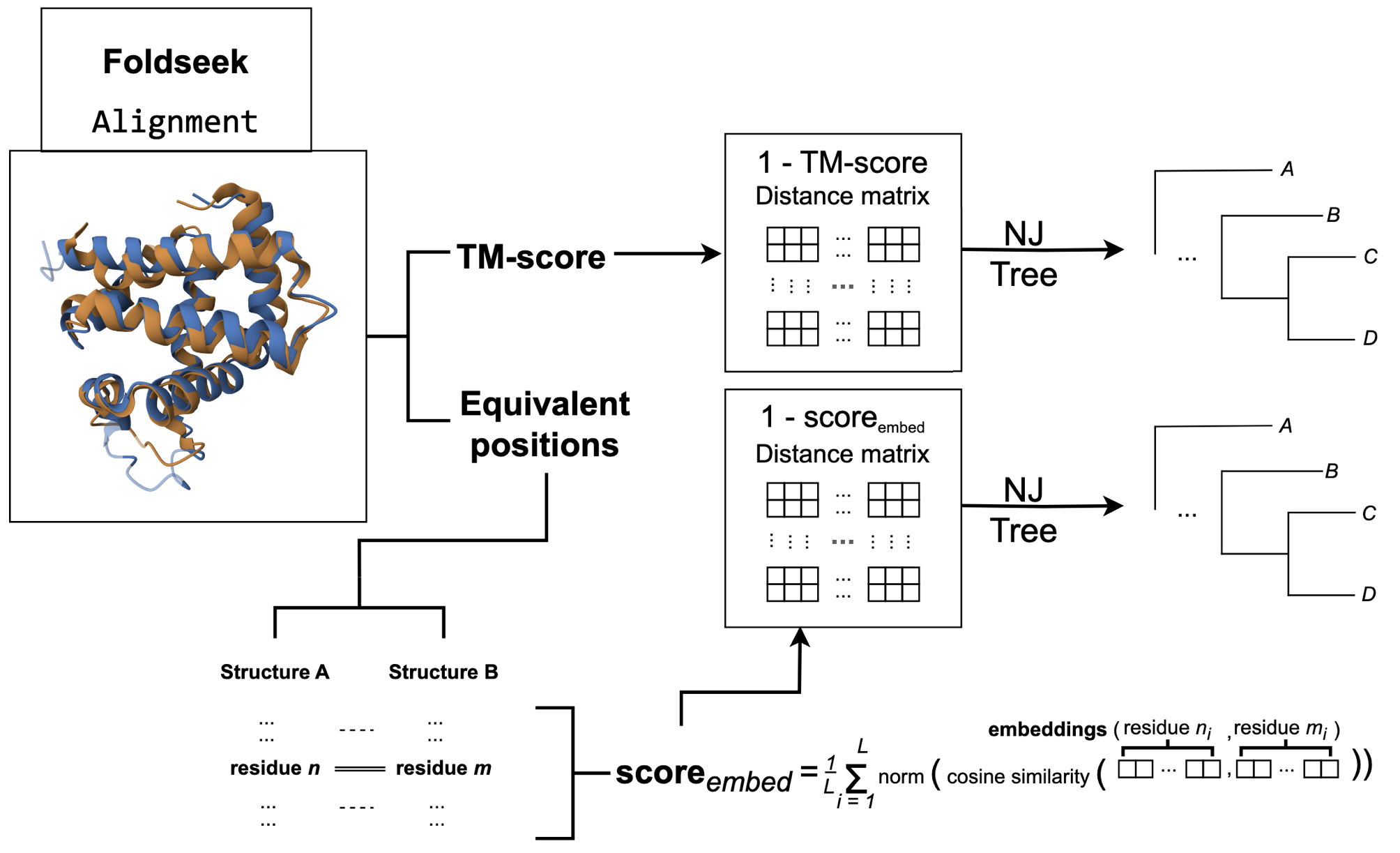
If moving from 1D sequences to 3D structures allows us to see further back in evolutionary time, can increasing the dimensionality of our data reveal these details better? We present Structome-DeepRoots, an exploration into whether a rich phylogenetic signal is an emergent property of high-dimensional protein representations.
Recent advances in deep learning have produced models capable of translating complex protein features into powerful, abstract high-dimensional latent spaces. DeepRoots taps into this abstracted space to generate evolutionary insights. We are excited to share that Structome-DeepRoots, shows a strong signal that this exploration has potential. This is our first attempt at digging through these abstract spaces to uncover the deep past.